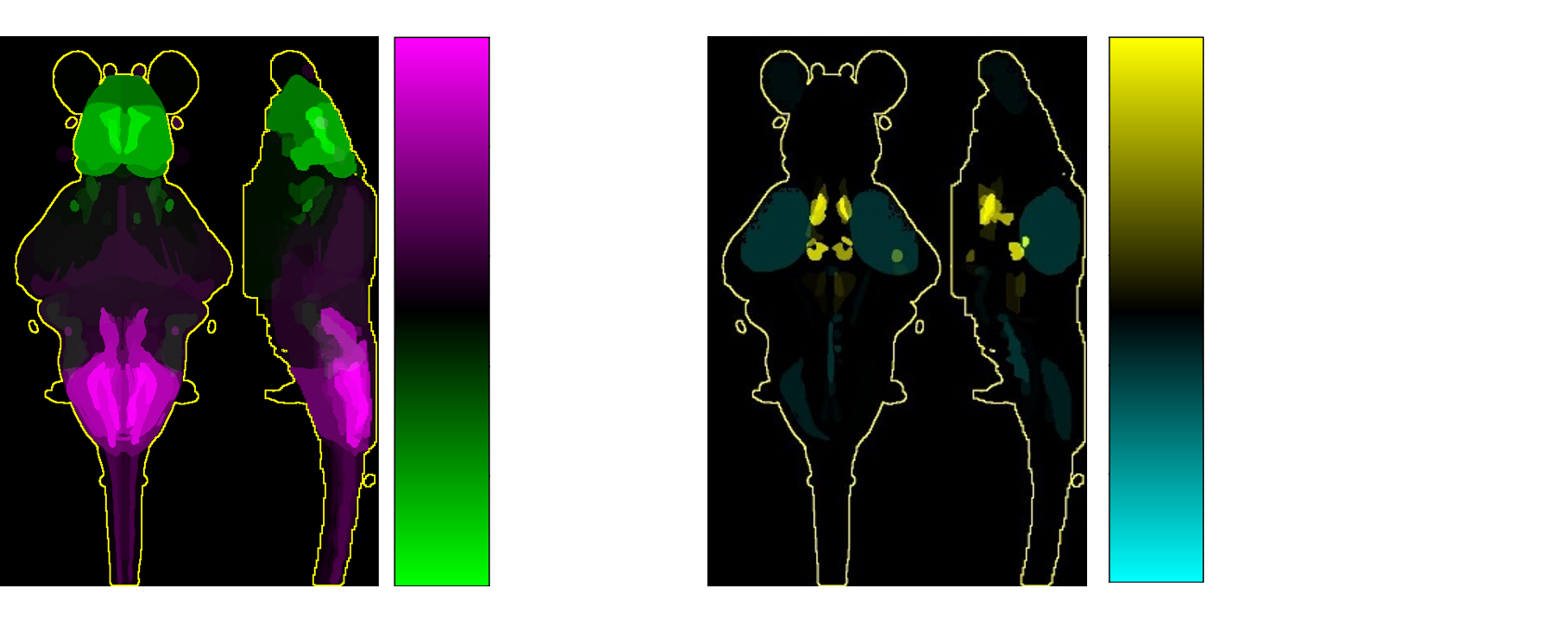In the ENHANCE lab
We are interested in advancing the understanding and treatment of diseases through equitable genomics with a specific focus on non-coding genetic elemnts. Leveraging human genomics data, we use zebrafish as a model to study genes and enhancers (an example of a non-coding element) in JAZF1 and metabolic disorders and the polygenic contribution to polycystic ovary syndrome. We use state-of-the-art molecular and cell biology techniques, including pILGET-mediated transgenesis whole-brain mapping, high-throughput behavioural phenotyping, luciferase assay, CRISPR gene editing and tail wounding, in dissecting the role of genetic elements of interest in health and disease.
Projects
Equitable Genomics - Advancing Research, Health, and Capability
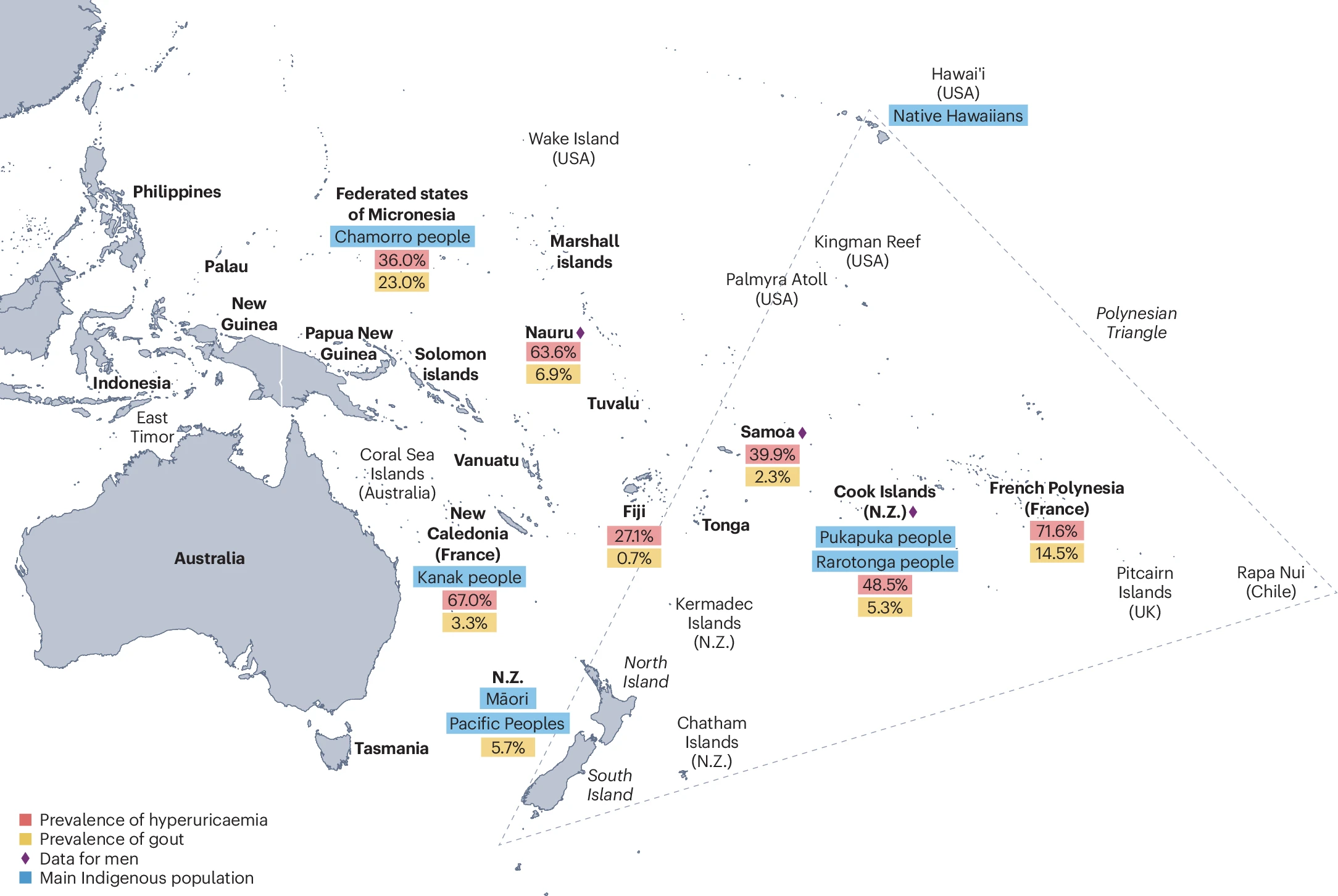
The ENHANCE lab is dedicated to understanding the genetic factors that influence health outcomes for Māori and Pasifika peoples. Many diseases, such as gout, are prevalent in these populations and have well-established underlying genetic components. Previous research has identified Māori- and Pacific-specific genetic variants linked to disease risk, and we aim to expand this knowledge by identifying new population-specific genetic variants and conducting functional analyses to uncover their biological effects. Genomic databases largely exclude Māori and Pasifika peoples, threatening to widen health inequities as precision medicine advances. By addressing this gap, our research will not only enhance the genomic representation of these populations in health-related studies but also uncover novel disease mechanisms relevant to all populations. Our findings could improve diagnostics, enable early interventions, and lead to more inclusive and effective treatments for all. Here in Aotearoa, we have a unique opportunity to explore how inherited genetic variation in these populations contributes to disease risk, ensuring Māori and Pasifika communities benefit equally from future medical innovations. As a Māori-led lab, we are also committed to strengthening Māori and Pacific genomics by recruiting and mentoring Māori and Pasifika students in this critical field while fostering strong relationships with the communities we serve. By elevating Māori and Pacific genomics into global research, we aim to create to a future where precision medicine is equitable and effective for all.
Non-Coding Genetic Elements
Non-coding elements are far more than just “junk” DNA. Genome-wide association studies indicate that >90% of variants associated with complex disease lie within the non-coding genome. Despite their significance in disease, there has been limited research on these elements. In contrast to the protein coding genome, where a variant will have more obvious functional consequences, these elements are a lot more complex and it is harder to decipher their effect on genes. These elements, such as enhancers, promoters, and silencers, are key regulators of gene expression. Promoters are involved in the initiation of transcription. Enhancers bind transcription factors that help drive gene expression whereas silencers act to repress it. Variants in these non-coding elements can greatly alter transcription which may contribute to distinct phenotypes. A classic example of this variation is when there is a change in the HERC2 enhancer sequence that causes decreased expression of the OCA2 gene, leading to an individual having blue eyes. Our lab investigates the contribution of the non-coding genome to metabolic conditions such as gout, diabetes, and polycystic ovary syndrome. These are all prevalent chronic conditions in which the aetiology is not well understood. Using zebrafish as a model organism, significant non-coding variants associated with these conditions can be investigated to understand their impact on gene expression.
JAZF1 and metabolic disorders
The development of complex diseases such as type two diabetes is multifactorial and involves interplay between both genetic and environmental factors. An approach known as Genome-Wide Association Studies (GWAS) is an important method to identify regions of the genome that associate with disease. Working with genomes from Māori and Pacific peoples has enabled the identification of unique, population-specific genetic elements that associate with metabolic conditions. Out of the many genetic elements identified in this analysis our project focuses on a variant that is found in a regulatory region of JAZF1. The JAZF1 gene acts in a number of metabolically active tissues (pancreas, brain, liver, muscle) to regulate energy balance. Of note, the JAZF1 gene encodes a transcription factor known to act in pancreatic beta-cells, regulating insulin secretion. Its role in other tissues such as the brain and liver is an emerging topic of study. Zebrafish are a powerful model organism to study gene function. Zebrafish express an orthologue of human JAZF1 known as jazf1b. The ENHANCE lab has taken the (human) JAZF1 genetic element coupled to green-fluorescent protein to visualise where the region is actively driving transcription in zebrafish. Preliminary data indicates extra-pancreatic function of JAZF1 and suggests a neuronal role including the brain’s control of circadian rhythm. To explore the behavioural, metabolic, and genetic consequences altering jazf1b in zebrafish, we generated a viable jazf1b knock out zebrafish strain. Our data thus far suggests that jazf1b is involved in circadian control supporting a neuronal function. Additionally, an elevated deposition of fat in the liver may drive JAZF1-mediated pathology. Our research aims to provide population-focused insights into the genetic role of JAZF1 and explore whether environmental stressors (elevated glucose or lipid exposure) worsen these effects.
Polycystic ovary syndrome
Polycystic ovary syndrome (PCOS) is the predominant cause of infertility worldwide and is extremely common here in Aotearoa, affecting ~5-15% of women of reproductive age. PCOS is characterized by irregular or reduced ovulation, high androgen hormone levels, and presence of multiple cysts on the ovaries. Despite its prevalence worldwide little is known about its underlying biological mechanisms.
Previous large genome-wide association studies (GWAS) of genetic determinants of PCOS have resulted in the identification of numerous loci. In the ENHANCE lab we seek to expand on this genetic information. Using PCOS GWAS data and expression quantitative trait loci (eQTL) data from GTEx we have found probable causal genes with genetic expression differences in the brain and ovaries. We are generating knockout models of these genes in zebrafish and the testing their contribution to the development of PCOS. We aim to identify novel potential therapeutic targets for the treatment of PCOS here in Aotearoa.
Techniques
MAPmap
We leverage zebrafish larvae transparency, smaller brain size and ease of genetic modification to perform high-throughput functional and structural mapping. We use MAPmap, a mitogen-activated protein (MAP) kinase-based whole-brain staining and imaging protocol to visualise and compare brain activation and region-specific changes in volume between different experimental groups. The readout allows us to identify brain areas that may be over or under -active, and detect deviations in brain formation or signs of neurodegeneration. These changes can be checked against a comprehensive reference atlas containing spatially resolved gene expression, cell type marker and protein expression data to further refine our understanding of affected brain circuits in disease.
High-throughput behavioural phenotyping
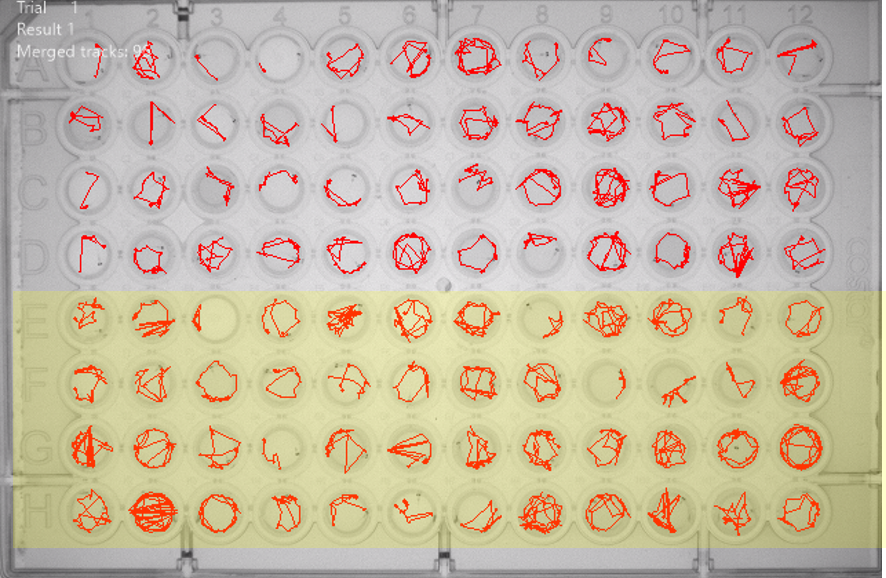
Behavioural analysis is crucial in understanding how our genetic models affect the physiology and brain function of larval zebrafish. We use a commercially available high-throughout solution (DanioVision, Noldus) that allows the simultaneous tracking of 96 fish across multiple days. In the system, we are able to alter ambient temperature, light intensity/duration in different colours and administer mechanical “taps” as a startle stimulus. We can combine this repertoire of manipulations to detect changes in circadian rhythms, visual/acoustic startle responses and habituation, sensory perception and locomotor functions. Changes in baseline and stimulus-bound movement trajectories offer us a valuable readout on how the brain interacts with the environment, providing insights to potential brain circuit dysfunction.
Tail wounding
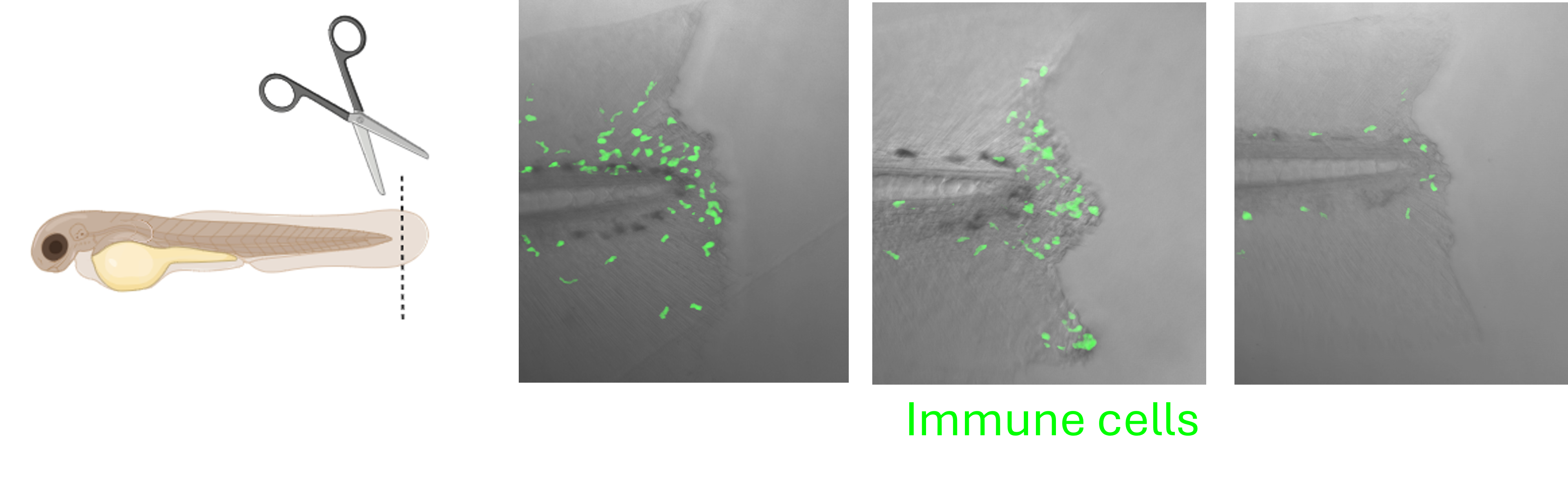
The immune system plays a key role in the daily maintenance and repair of the body during health and disease and is pivotal for protection against invading microorganisms such as bacteria. The cells of the immune system (‘white blood cells’) carry out numerous dynamic activities in response to stimulation including infection and injury. Model animals, including zebrafish, enable the study of immune cell behaviour. Zebrafish have a functional innate immune system from early embryonic stages and into adulthood. Many haematopoietic (blood) pathways that underly an immune response are similar in zebrafish as in humans. Zebrafish larvae are transparent and through genetic editing can produce fluorescent immune cells to facilitate immune-focused research.
In our lab, we use a simple larval tailfin wound to stimulate innate immune cells (macrophages and neutrophils) into action. The larvae that we wound are typically deficient in a gene that might impact immune processes or have been treated with potential immune altering substances (such as glucose). We are interested in any changes to immune cell behaviour under our experimental conditions. The speed of recruitment to the wound, how long the cells stay at the injury, and how many cells enter the wound site are all important parameters we can measure to understand how the immune cells (and therefore the wider immune system) are functioning in our models.
pIGLET-mediated transgenesis
PhiC31 Integrase Genomic Loci Engineered for Transgenesis (pIGLET) is a recently developed zebrafish transgenesis system that we use in our lab to model human non-coding regulatory elements. As the name suggests, pIGLET refers to a transgenic zebrafish line containing a unique attP landing site located within a well-mapped genomic region that avoids both phenotypic consequences and positional effects. We utilize two lines of pIGLET fish: pIGLET14a and pIGLET24b, with the end numbers indicating the chromosomal location of the landing site. These lines were generously provided in collaboration with Dr. Robert Lalonde and Professor Christian Mosimann at the University of Colorado Anschutz. By injecting a plasmid containing complementary attB sites flanking a region of interest, alongside phiC31 integrase, we achieve single-site, unidirectional integration of that region into the genome at the designated site. This enables us to generate transgenic lines housing putative enhancer regions positioned upstream of a fluorescent reporter. Doing so facilitates both tissue-specific expression profiling and in vivo quantification of enhancer activity across populations, capabilities that are limited or absent in other zebrafish transgenesis methods. Through this system, we are able to investigate the effects of variation within enhancer regions, with the ultimate goal of translating our findings into improved health outcomes.
Luciferase assay
Reporter assays, such as the luciferase assay, are one of the key tools we use to investigate how non-coding genetic variants influence disease risk. This method harnesses the luciferase gene which encodes a bioluminescent enzyme originally identified in fireflies. By linking suspected regulatory DNA sequences, including those with or without genetic variants, to a luciferase reporter gene and introducing them into human cell models, such as liver, kidney, neuronal, or immune cells, we can measure changes in luminescence as a direct readout of regulatory activity. If a variant alters this luminescence it suggests that the variant influences how and when downstream gene expression is switched on or off. We apply this technique to study non-coding variants associated with diseases such as gout, diabetes, and PCOS, providing crucial insights into how changes in gene regulation contribute to complex disease biology.
CRISPR gene editing
CRISPR-Cas9 is a powerful and precise gene-editing technology that enables targeted modifications to DNA within living organisms. Originally derived from an adaptive immune system found in bacteria, the CRISPR-Cas9 system uses a guide RNA (gRNA) complementary to the target DNA sequence to recruit the Cas9 enzyme, which introduces a double-stranded break at the specific site. This break is then repaired by the cell’s machinery via either non-homologous end joining (NHEJ), often resulting in insertions or deletions (indels), or homology-directed repair (HDR), where a donor DNA template with homology arms enables precise sequence insertion. Recent advancements have expanded CRISPR-Cas technologies to include CRISPRa/i (activation/interference), which uses a deactivated Cas9 (dCas9) fused to effector molecules to upregulate or repress gene expression, and base editing, which employs dCas9 coupled with a deaminase enzyme to directly convert one DNA base into another without inducing double-stranded breaks. Here in the ENHANCE laboratory, we harness these CRISPR-Cas9-based methodologies to introduce single-nucleotide changes and knock out genes in cellular models and zebrafish. Our research focuses on advancing the understanding and treatment of diseases through equitable genomics, with a particular emphasis on non-coding genetic elements. We investigate the genetic contributions to complex diseases such as gout, diabetes, and polycystic ovary syndrome (PCOS), with the ultimate goal of developing robust disease models to drive therapeutic discovery.